Figs. S3 - S14
NOTE: Run all the code included on the “Base Code” page prior to running the following.
Setting Up
Load eigenworms and worm data of choice.
# working directory
setwd("") # enter appropriate working directory
# load eigenworms [Broekmans et al. 2016]
ew = read.csv("eigenworms.csv", header=F, sep=",")
# load coefficients from worm of choice and crop the data to specified values
start = 10001
end = 12000
data = read.csv("12 Foraging Worms/1.txt", header=F, sep="")[start:end,1:5]Variables
These are the optimal parameters for the data produced by Broekmans et al. 2016.
E = 5 # embedding dimension
tp = 1
theta = 2 # linearity
tau = 1
# library and prediction selection
lib = c(1,1000)
pred = c(1,2000)
# convert frames to seconds (i.e. indicate frames per second as fps)
fps = 16 # foraging wormsRun Functions
Run the functions created on the “Base Code” page.
# run the embedding function
matrix <- make_embed(data, E, tau, tp)
# run the prediction function
new_pred <- make_pred(E, matrix, theta, lib, pred)
observations_total = new_pred$obs
predictions_total = new_pred$pred
# run the EDM error function
errors <- edm_error(pred, observations_total, ew, predictions_total)
# run the data classification function
new_class <- data_class(data, pred)
forward = new_class$forw
backward = new_class$back
turns = new_class$turn
change = new_class$chanError Plot
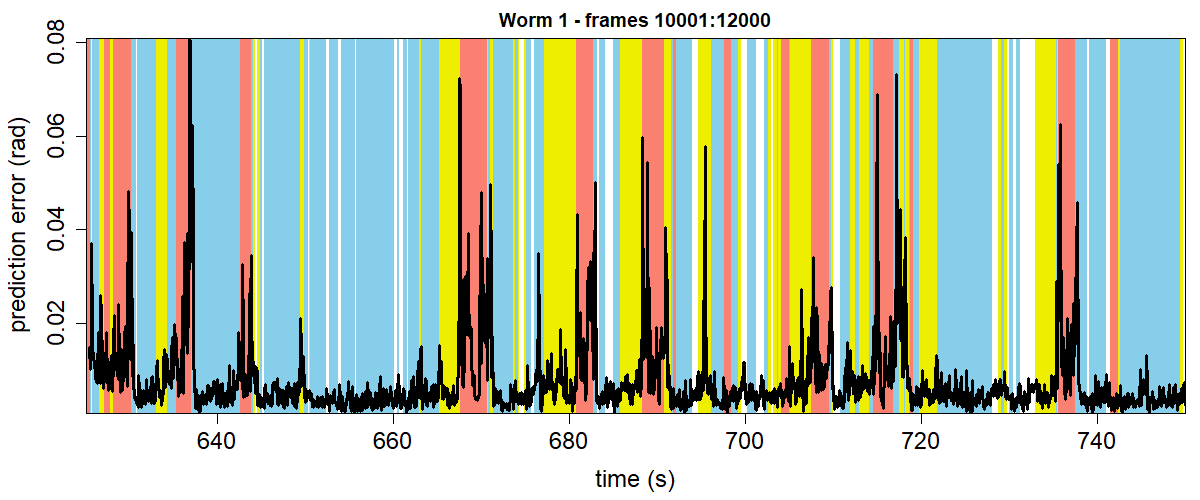
Create an RMS error vs time plot with colored classifications.
# setup the RMS error vs time plot
par(mai=c(0.9,0.9,0.4,0.15))
plot(c(start:end)/fps, errors, col="white", xlab="time (s)", ylab="prediction error (rad)",
cex.lab=1.5, cex.axis=1.5, xaxs="i", yaxs="i", main="Worm 1 - frames 10001:12000")
# add colored rectangles for forwards, backwards, and turns
for (i in 1:length(forward)){
rect(c(start:end)[forward[i]]/fps, min(errors, na.rm=TRUE),
c(start:end)[forward[i]+2]/fps, max(errors, na.rm=TRUE),
col="skyblue", border=NA)
}
for (i in 1:length(backward)){
rect(c(start:end)[backward[i]]/fps, min(errors, na.rm=TRUE),
c(start:end)[backward[i]+2]/fps, max(errors, na.rm=TRUE),
col="yellow2", border=NA)
}
for (i in 1:length(turns)){
rect(c(start:end)[turns[i]]/fps, min(errors, na.rm=TRUE),
c(start:end)[turns[i]+2]/fps, max(errors, na.rm=TRUE),
col="salmon", border=NA)
}
# add the error line on top
lines(c(start:end)/fps, errors, lwd=3)
box(col="black")